bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
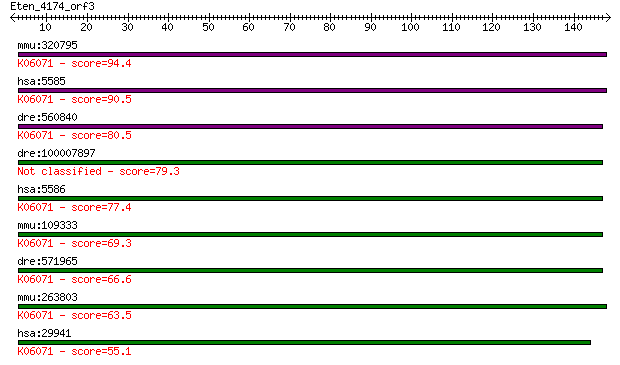
Query= Eten_4174_orf3
Length=148
Score E
Sequences producing significant alignments: (Bits) Value
mmu:320795 Pkn1, DBK, F730027O18Rik, PAK1, PRK1, Pkn, Prkcl1, ... 94.4 1e-19
hsa:5585 PKN1, DBK, MGC46204, PAK-1, PAK1, PKN, PKN-ALPHA, PRK... 90.5 2e-18
dre:560840 hypothetical LOC560840; K06071 protein kinase N [EC... 80.5 2e-15
dre:100007897 MGC162290; zgc:162290 79.3 4e-15
hsa:5586 PKN2, MGC150606, MGC71074, PAK2, PRK2, PRKCL2, PRO204... 77.4 2e-14
mmu:109333 Pkn2, 6030436C20Rik, AI507382, PRK2, Prkcl2, Stk7; ... 69.3 4e-12
dre:571965 protein kinase N2-like; K06071 protein kinase N [EC... 66.6 3e-11
mmu:263803 Pkn3, AW209115, BC034126, MGC31699; protein kinase ... 63.5 2e-10
hsa:29941 PKN3, RP11-545E17.1; protein kinase N3 (EC:2.7.11.13... 55.1 7e-08
> mmu:320795 Pkn1, DBK, F730027O18Rik, PAK1, PRK1, Pkn, Prkcl1,
Stk3; protein kinase N1 (EC:2.7.11.13); K06071 protein kinase
N [EC:2.7.11.13]
Length=951
Score = 94.4 bits (233), Expect = 1e-19, Method: Compositional matrix adjust.
Identities = 52/145 (35%), Positives = 63/145 (43%), Gaps = 0/145 (0%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF V +P F IG +L NP
Sbjct 806 DWWGLGVLLYEMLVGESPFPGDDEEEVFDSIVNDEVRYPRFLSAEAIGIMRRLLRRNPER 865
Query 63 GLGPPGGGFGGGKKPPFFRALGGGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPGG 122
LG KK PFFR+LG L+ R PP F P G +F F G AP
Sbjct 866 RLGSTERDAEDVKKQPFFRSLGWDVLLARRLPPPFVPTLSGRTDVSNFDEEFTGEAPTLS 925
Query 123 PPRGGGPLPPPKKGAFRGFNFGGGG 147
PPR PL ++ AFR F+F GG
Sbjct 926 PPRDARPLTAAEQAAFRDFDFVAGG 950
> hsa:5585 PKN1, DBK, MGC46204, PAK-1, PAK1, PKN, PKN-ALPHA, PRK1,
PRKCL1; protein kinase N1 (EC:2.7.11.13); K06071 protein
kinase N [EC:2.7.11.13]
Length=942
Score = 90.5 bits (223), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 51/145 (35%), Positives = 62/145 (42%), Gaps = 0/145 (0%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF V +P F IG +L NP
Sbjct 797 DWWGLGVLLYEMLVGESPFPGDDEEEVFDSIVNDEVRYPRFLSAEAIGIMRRLLRRNPER 856
Query 63 GLGPPGGGFGGGKKPPFFRALGGGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPGG 122
LG KK PFFR LG +L+ R PP F P G +F F G AP
Sbjct 857 RLGSSERDAEDVKKQPFFRTLGWEALLARRLPPPFVPTLSGRTDVSNFDEEFTGEAPTLS 916
Query 123 PPRGGGPLPPPKKGAFRGFNFGGGG 147
PPR PL ++ AF F+F GG
Sbjct 917 PPRDARPLTAAEQAAFLDFDFVAGG 941
> dre:560840 hypothetical LOC560840; K06071 protein kinase N [EC:2.7.11.13]
Length=948
Score = 80.5 bits (197), Expect = 2e-15, Method: Compositional matrix adjust.
Identities = 45/144 (31%), Positives = 58/144 (40%), Gaps = 0/144 (0%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF V +P F IG +L NP
Sbjct 803 DWWGLGVLIYEMLVGESPFPGDDEEEVFDSIVNDEVRYPRFLSAEAIGIMRRLLRRNPER 862
Query 63 GLGPPGGGFGGGKKPPFFRALGGGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPGG 122
LG KK PFFR + +L+ R PP F P GG +F F AP
Sbjct 863 RLGSSEKDAEDVKKQPFFRNMDWEALLLRKLPPPFIPSIGGKEDVSNFDEEFTTEAPTLT 922
Query 123 PPRGGGPLPPPKKGAFRGFNFGGG 146
PPR L + +FR F++
Sbjct 923 PPREPRVLSRKDQDSFRDFDYVSD 946
> dre:100007897 MGC162290; zgc:162290
Length=909
Score = 79.3 bits (194), Expect = 4e-15, Method: Compositional matrix adjust.
Identities = 42/144 (29%), Positives = 55/144 (38%), Gaps = 0/144 (0%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF V +P F IG +L NP
Sbjct 764 DWWGLGVLIYEMMVGESPFPGDDEEEVFDSIVNDEVRYPRFLSTEAIGIMRRLLRRNPER 823
Query 63 GLGPPGGGFGGGKKPPFFRALGGGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPGG 122
LG KK PFFR + +L+ R PP F P +F F AP
Sbjct 824 RLGSGEKDAEDIKKQPFFRNMDWDALLQRKVPPPFVPAVANSEDVSNFDAEFTNEAPTLT 883
Query 123 PPRGGGPLPPPKKGAFRGFNFGGG 146
PPR L + F+ F++
Sbjct 884 PPRERRSLSRKDQDYFKDFDYVSD 907
> hsa:5586 PKN2, MGC150606, MGC71074, PAK2, PRK2, PRKCL2, PRO2042,
Pak-2; protein kinase N2 (EC:2.7.11.13); K06071 protein
kinase N [EC:2.7.11.13]
Length=984
Score = 77.4 bits (189), Expect = 2e-14, Method: Compositional matrix adjust.
Identities = 41/144 (28%), Positives = 55/144 (38%), Gaps = 0/144 (0%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF V +P F I +L NP
Sbjct 839 DWWGLGVLIYEMLVGESPFPGDDEEEVFDSIVNDEVRYPRFLSTEAISIMRRLLRRNPER 898
Query 63 GLGPPGGGFGGGKKPPFFRALGGGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPGG 122
LG KK PFFR + +L+ + P F P G +F F AP
Sbjct 899 RLGASEKDAEDVKKHPFFRLIDWSALMDKKVKPPFIPTIRGREDVSNFDDEFTSEAPILT 958
Query 123 PPRGGGPLPPPKKGAFRGFNFGGG 146
PPR L ++ FR F++
Sbjct 959 PPREPRILSEEEQEMFRDFDYIAD 982
> mmu:109333 Pkn2, 6030436C20Rik, AI507382, PRK2, Prkcl2, Stk7;
protein kinase N2 (EC:2.7.11.13); K06071 protein kinase N
[EC:2.7.11.13]
Length=983
Score = 69.3 bits (168), Expect = 4e-12, Method: Compositional matrix adjust.
Identities = 40/144 (27%), Positives = 53/144 (36%), Gaps = 0/144 (0%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF V +P F I +L NP
Sbjct 838 DWWGLGVLIYEMLVGESPFPGDDEEEVFDSIVNDEVRYPRFLSTEAISIMRRLLRRNPER 897
Query 63 GLGPPGGGFGGGKKPPFFRALGGGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPGG 122
LG KK PFFR +L+ + P F P G +F F AP
Sbjct 898 RLGAGEKDAEDVKKHPFFRLTDWSALMDKKVKPPFVPTIRGREDVSNFDDEFTSEAPILT 957
Query 123 PPRGGGPLPPPKKGAFRGFNFGGG 146
PPR L ++ F F++
Sbjct 958 PPREPRILLEEEQEMFHDFDYVAD 981
> dre:571965 protein kinase N2-like; K06071 protein kinase N [EC:2.7.11.13]
Length=970
Score = 66.6 bits (161), Expect = 3e-11, Method: Compositional matrix adjust.
Identities = 40/145 (27%), Positives = 52/145 (35%), Gaps = 2/145 (1%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF V +P I +L NP
Sbjct 825 DWWGLGVLIFEMLVGESPFPGDDEEEVFDSIVNDEVRYPRVLSTEAISIMRRLLRRNPER 884
Query 63 GLGPPGGGFGGGKKPPFFRALG-GGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPG 121
LG KK FFR + G L + PP F P +F F AP
Sbjct 885 RLGAAERDAEDVKKHLFFRDIDWDGLLAKKVKPP-FVPTIQCSSDVSNFDDEFTSEAPVL 943
Query 122 GPPRGGGPLPPPKKGAFRGFNFGGG 146
PPR PL ++ F F++
Sbjct 944 TPPREPRPLNQHEQDLFTDFDYIAD 968
> mmu:263803 Pkn3, AW209115, BC034126, MGC31699; protein kinase
N3 (EC:2.7.11.13); K06071 protein kinase N [EC:2.7.11.13]
Length=878
Score = 63.5 bits (153), Expect = 2e-10, Method: Compositional matrix adjust.
Identities = 41/145 (28%), Positives = 53/145 (36%), Gaps = 0/145 (0%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF V P F + +L +P
Sbjct 730 DWWGLGVLLYEMLVGECPFPGDTEEEVFECIVSADVPCPHFLSVQGLELIQKLLQKSPEK 789
Query 63 GLGPPGGGFGGGKKPPFFRALGGGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPGG 122
LG K PFFR +L+ R P F P GP +F G F P
Sbjct 790 RLGAGERDAEEIKVQPFFRTTNWQALLARTVQPPFVPTLCGPADLRYFEGEFTSLPPTLT 849
Query 123 PPRGGGPLPPPKKGAFRGFNFGGGG 147
PP L ++ AFR F+F
Sbjct 850 PPVSQSSLTARQQAAFRDFDFVSEQ 874
> hsa:29941 PKN3, RP11-545E17.1; protein kinase N3 (EC:2.7.11.13);
K06071 protein kinase N [EC:2.7.11.13]
Length=889
Score = 55.1 bits (131), Expect = 7e-08, Method: Compositional matrix adjust.
Identities = 42/141 (29%), Positives = 54/141 (38%), Gaps = 0/141 (0%)
Query 3 EWGGPGLFFLKKGGGGPPFPGGWGGGVFSPFFQGGVFFPPFFLRGTIGFFAGVLGGNPGG 62
+W G G+ + G PFPG VF +P F + F +L P
Sbjct 741 DWWGLGVLLYEMLVGECPFPGDTEEEVFDCIVNMDAPYPGFLSVQGLEFIQKLLQKCPEK 800
Query 63 GLGPPGGGFGGGKKPPFFRALGGGSLVGRAPPPLFFPPGGGPPCFPHFWGGFPGGAPPGG 122
LG K PFFR +L+ R P F P GP +F G F G P
Sbjct 801 RLGAGEQDAEEIKVQPFFRTTNWQALLARTIQPPFVPTLCGPADLRYFEGEFTGLPPALT 860
Query 123 PPRGGGPLPPPKKGAFRGFNF 143
PP L ++ AFR F+F
Sbjct 861 PPAPHSLLTARQQAAFRDFDF 881
Lambda K H
0.323 0.163 0.605
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3003468616
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40