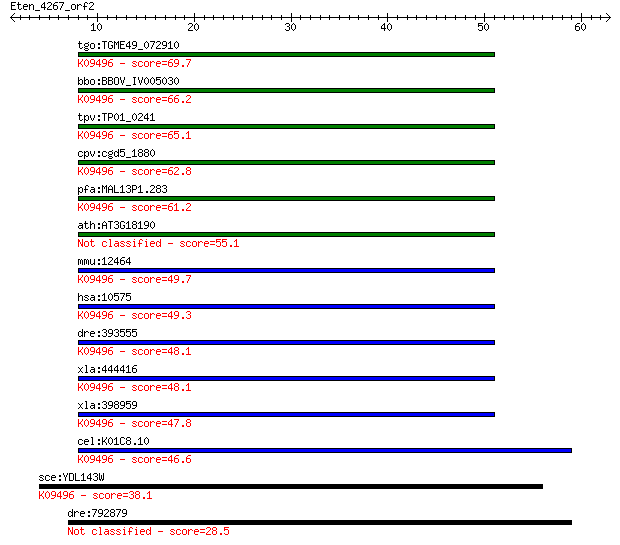
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_4267_orf2
Length=62
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_072910 TCP-1/cpn60 family chaperonin, putative ; K0... 69.7 2e-12
bbo:BBOV_IV005030 23.m06469; T-complex protein 1 delta subunit... 66.2 2e-11
tpv:TP01_0241 chaperonin 60 kDa; K09496 T-complex protein 1 su... 65.1 5e-11
cpv:cgd5_1880 conserved probable chaperonin containing TCP-1 d... 62.8 3e-10
pfa:MAL13P1.283 TCP-1/cpn60 chaperonin family, putative; K0949... 61.2 8e-10
ath:AT3G18190 chaperonin, putative 55.1 5e-08
mmu:12464 Cct4, 2610204B21Rik, A45, C78323, Cctd; chaperonin c... 49.7 2e-06
hsa:10575 CCT4, CCT-DELTA, Cctd, MGC126164, MGC126165, SRB; ch... 49.3 3e-06
dre:393555 cct4, MGC65789, zgc:65789, zgc:77192; chaperonin co... 48.1 6e-06
xla:444416 cct4, MGC82994; chaperonin containing TCP1, subunit... 48.1 7e-06
xla:398959 MGC83370; hypothetical protein LOC398959; K09496 T-... 47.8 8e-06
cel:K01C8.10 cct-4; Chaperonin Containing TCP-1 family member ... 46.6 2e-05
sce:YDL143W CCT4, ANC2, TCP4; Cct4p; K09496 T-complex protein ... 38.1 0.007
dre:792879 hypothetical LOC792879 28.5 5.6
> tgo:TGME49_072910 TCP-1/cpn60 family chaperonin, putative ;
K09496 T-complex protein 1 subunit delta
Length=542
Score = 69.7 bits (169), Expect = 2e-12, Method: Composition-based stats.
Identities = 35/43 (81%), Positives = 37/43 (86%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
RMLRE+R+LIAKM K IAAT C LLIQKSILRDAV DLSLDY
Sbjct 278 RMLREERILIAKMVKQIAATGCNVLLIQKSILRDAVNDLSLDY 320
> bbo:BBOV_IV005030 23.m06469; T-complex protein 1 delta subunit;
K09496 T-complex protein 1 subunit delta
Length=535
Score = 66.2 bits (160), Expect = 2e-11, Method: Composition-based stats.
Identities = 32/43 (74%), Positives = 37/43 (86%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R++RE+RL+ A+M K IAAT C LLIQKSILRDAVTDLSLDY
Sbjct 271 RIMREERLITARMVKQIAATGCNVLLIQKSILRDAVTDLSLDY 313
> tpv:TP01_0241 chaperonin 60 kDa; K09496 T-complex protein 1
subunit delta
Length=531
Score = 65.1 bits (157), Expect = 5e-11, Method: Composition-based stats.
Identities = 32/43 (74%), Positives = 36/43 (83%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R++RE+RL+ AKM K IA T C LLIQKSILRDAVTDLSLDY
Sbjct 268 RIMREERLITAKMVKKIAQTGCNVLLIQKSILRDAVTDLSLDY 310
> cpv:cgd5_1880 conserved probable chaperonin containing TCP-1
delta subunit ; K09496 T-complex protein 1 subunit delta
Length=550
Score = 62.8 bits (151), Expect = 3e-10, Method: Compositional matrix adjust.
Identities = 31/43 (72%), Positives = 37/43 (86%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R+LRE+RLLIAKM K IAAT C LLIQKSILR+A++ L+LDY
Sbjct 286 RLLREERLLIAKMIKQIAATGCNVLLIQKSILREAISTLALDY 328
> pfa:MAL13P1.283 TCP-1/cpn60 chaperonin family, putative; K09496
T-complex protein 1 subunit delta
Length=529
Score = 61.2 bits (147), Expect = 8e-10, Method: Composition-based stats.
Identities = 29/43 (67%), Positives = 36/43 (83%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R+LRE+RL+I KM K IA+T C L+IQKSILRDAV DL+LD+
Sbjct 266 RLLREERLIIGKMIKKIASTGCNLLIIQKSILRDAVNDLALDF 308
> ath:AT3G18190 chaperonin, putative
Length=536
Score = 55.1 bits (131), Expect = 5e-08, Method: Composition-based stats.
Identities = 29/43 (67%), Positives = 32/43 (74%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R+L+E+R I M K I AT C LLIQKSILRDAVTDLSL Y
Sbjct 272 RILKEERNYILGMIKKIKATGCNVLLIQKSILRDAVTDLSLHY 314
> mmu:12464 Cct4, 2610204B21Rik, A45, C78323, Cctd; chaperonin
containing Tcp1, subunit 4 (delta); K09496 T-complex protein
1 subunit delta
Length=539
Score = 49.7 bits (117), Expect = 2e-06, Method: Composition-based stats.
Identities = 24/43 (55%), Positives = 31/43 (72%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R+LRE+R I + K I T C LLIQKSILRDA++DL+L +
Sbjct 274 RVLREERAYILNLVKQIKKTGCNVLLIQKSILRDALSDLALHF 316
> hsa:10575 CCT4, CCT-DELTA, Cctd, MGC126164, MGC126165, SRB;
chaperonin containing TCP1, subunit 4 (delta); K09496 T-complex
protein 1 subunit delta
Length=539
Score = 49.3 bits (116), Expect = 3e-06, Method: Composition-based stats.
Identities = 24/43 (55%), Positives = 31/43 (72%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R+LRE+R I + K I T C LLIQKSILRDA++DL+L +
Sbjct 274 RVLREERAYILNLVKQIKKTGCNVLLIQKSILRDALSDLALHF 316
> dre:393555 cct4, MGC65789, zgc:65789, zgc:77192; chaperonin
containing TCP1, subunit 4 (delta); K09496 T-complex protein
1 subunit delta
Length=533
Score = 48.1 bits (113), Expect = 6e-06, Method: Composition-based stats.
Identities = 22/43 (51%), Positives = 30/43 (69%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R+LRE+R I + K + C LLIQKSILRDA++DL+L +
Sbjct 268 RVLREERAYILNLVKQVKKAGCTVLLIQKSILRDALSDLALHF 310
> xla:444416 cct4, MGC82994; chaperonin containing TCP1, subunit
4 (delta); K09496 T-complex protein 1 subunit delta
Length=539
Score = 48.1 bits (113), Expect = 7e-06, Method: Composition-based stats.
Identities = 23/43 (53%), Positives = 30/43 (69%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R+LRE+R I + K I C LLIQKSILRDA++DL+L +
Sbjct 274 RVLREERTYILNLVKQIKKVGCNVLLIQKSILRDALSDLALHF 316
> xla:398959 MGC83370; hypothetical protein LOC398959; K09496
T-complex protein 1 subunit delta
Length=541
Score = 47.8 bits (112), Expect = 8e-06, Method: Composition-based stats.
Identities = 23/43 (53%), Positives = 30/43 (69%), Gaps = 0/43 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDY 50
R+LRE+R I + K I C LLIQKSILRDA++DL+L +
Sbjct 276 RVLREERTYILNLVKQIKKAGCNVLLIQKSILRDALSDLALHF 318
> cel:K01C8.10 cct-4; Chaperonin Containing TCP-1 family member
(cct-4); K09496 T-complex protein 1 subunit delta
Length=540
Score = 46.6 bits (109), Expect = 2e-05, Method: Composition-based stats.
Identities = 23/51 (45%), Positives = 32/51 (62%), Gaps = 0/51 (0%)
Query 8 RMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDYRGESNNTC 58
R L+E+R + ++ K I A C LLIQKSILRDAV +L+L + + C
Sbjct 274 RALKEERQYLLEICKQIKAAGCNVLLIQKSILRDAVNELALHFLAKMKIMC 324
> sce:YDL143W CCT4, ANC2, TCP4; Cct4p; K09496 T-complex protein
1 subunit delta
Length=528
Score = 38.1 bits (87), Expect = 0.007, Method: Composition-based stats.
Identities = 23/52 (44%), Positives = 33/52 (63%), Gaps = 0/52 (0%)
Query 4 RKWHRMLREDRLLIAKMGKHIAATACICLLIQKSILRDAVTDLSLDYRGESN 55
R+ ++L+E+R + + K I C LLIQKSILRDAV DL+L + + N
Sbjct 258 RQMDKILKEERAYLLNICKKIKKAKCNVLLIQKSILRDAVNDLALHFLSKLN 309
> dre:792879 hypothetical LOC792879
Length=683
Score = 28.5 bits (62), Expect = 5.6, Method: Composition-based stats.
Identities = 20/54 (37%), Positives = 28/54 (51%), Gaps = 2/54 (3%)
Query 7 HRMLREDRLLIAKMG-KHIAATACICLLIQKSILRDAVTDLSLDYRGES-NNTC 58
HR+ + L I + +H+AA A +C L K D V L +RGES +TC
Sbjct 424 HRIGKRMSLSITEGALQHLAADALLCPLDSKLGFSDPVAQAVLHFRGESIADTC 477
Lambda K H
0.328 0.138 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2060478756
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40