bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_6626_orf3
Length=177
Score E
Sequences producing significant alignments: (Bits) Value
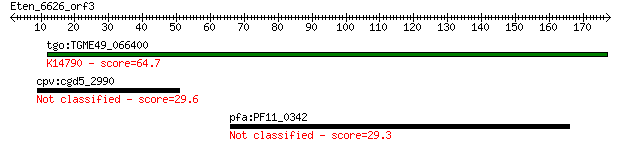
tgo:TGME49_066400 hypothetical protein ; K14790 nucleolar prot... 64.7 2e-10
cpv:cgd5_2990 hypothetical protein 29.6 6.1
pfa:PF11_0342 conserved Plasmodium protein 29.3 7.6
> tgo:TGME49_066400 hypothetical protein ; K14790 nucleolar protein
9
Length=960
Score = 64.7 bits (156), Expect = 2e-10, Method: Composition-based stats.
Identities = 51/176 (28%), Positives = 91/176 (51%), Gaps = 15/176 (8%)
Query 12 KNYTAYMKCEIYKFTKDKEEWQKRQEKRSKTRELFKDIISDAAIEVNNQEAADLLPDNPK 71
KNY AY+ CEI+ F +++E W RQEK+SKTRELFKD++++ + + A+ +
Sbjct 739 KNYAAYVNCEIHTFKRNEEGWATRQEKKSKTRELFKDLLAEEGDQAEDVVEAEKTEKHAA 798
Query 72 GKKAEA--ESTPLQNEGKGFRK---------PKNENKNKKGTINGDEEEEINTKGKAEKE 120
+ A A E + + G + + + + + +GD EE +T + KE
Sbjct 799 HEAARALLERDAVAQQLLGVDEKKKSKKTVEKRKKKQAPESDSDGDVFEESST---SRKE 855
Query 121 AKRAIDEIFLGAFEEKQKGKENKKKKQLTGKAKENSNPPAPSNENENANLNKTKNK 176
A+ A+D IF A +E QKGK + K + T A ++++ + ++ + + + K
Sbjct 856 AQEAVDTIFSEAAKE-QKGKRRRTKTEQTELADQDADGASAADSSSGTSKKRKGEK 910
> cpv:cgd5_2990 hypothetical protein
Length=764
Score = 29.6 bits (65), Expect = 6.1, Method: Compositional matrix adjust.
Identities = 17/42 (40%), Positives = 24/42 (57%), Gaps = 1/42 (2%)
Query 9 LREKNYTAYMKCEIYKFTKDKEEWQKRQEKRSKTRELFKDII 50
L KN+ + +I F K K++W EK KTR+LF +II
Sbjct 660 LMSKNFKIFKSLKIESFNK-KDDWNNVNEKIDKTRKLFSNII 700
> pfa:PF11_0342 conserved Plasmodium protein
Length=2072
Score = 29.3 bits (64), Expect = 7.6, Method: Compositional matrix adjust.
Identities = 30/100 (30%), Positives = 48/100 (48%), Gaps = 3/100 (3%)
Query 66 LPDNPKGKKAEAESTPLQNEGKGFRKPKNENKNKKGTINGDEEEEINTKGKAEKEAKRAI 125
+ D+ + + TP +N+GK KN + + ++ +N G+ +K+ R I
Sbjct 307 ILDDSSNIQFHKKETPSKNQGKNKYIFKNVERKNTAVYDKNKNTSLNNYGQQQKKINR-I 365
Query 126 DEIFLGAFEEKQKGKENKKKKQLTGKAKENSNPPAPSNEN 165
DEIF EK+K KE ++K L K N N +NEN
Sbjct 366 DEIFKNLI-EKRKKKEMRQKYMLKKKFLMN-NYKGKTNEN 403
Lambda K H
0.301 0.123 0.330
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4665550176
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40