bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
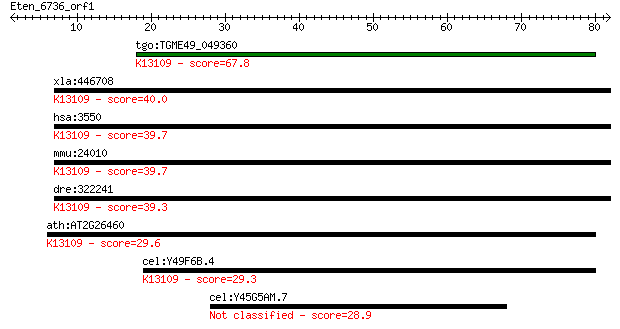
Query= Eten_6736_orf1
Length=81
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_049360 hypothetical protein ; K13109 IK cytokine 67.8 9e-12
xla:446708 MGC83922 protein; K13109 IK cytokine 40.0 0.002
hsa:3550 IK, CSA2, MGC59741, RED; IK cytokine, down-regulator ... 39.7 0.002
mmu:24010 Ik, MuRED; IK cytokine; K13109 IK cytokine 39.7 0.003
dre:322241 ik, wu:fb54c03, zgc:56500; IK cytokine; K13109 IK c... 39.3 0.003
ath:AT2G26460 RED family protein; K13109 IK cytokine 29.6 2.4
cel:Y49F6B.4 smu-2; Suppressor of Mec and Unc defects family m... 29.3 3.4
cel:Y45G5AM.7 hypothetical protein 28.9 4.3
> tgo:TGME49_049360 hypothetical protein ; K13109 IK cytokine
Length=586
Score = 67.8 bits (164), Expect = 9e-12, Method: Compositional matrix adjust.
Identities = 34/62 (54%), Positives = 45/62 (72%), Gaps = 1/62 (1%)
Query 18 QQFASKAAYEEYMNSIELVPKAAYSYGRRIAGDINSSKNKKKQKLNPEQEWRAIEKLIAE 77
QQF SK YE+Y+ IE VPKAA YGRR+AG+ + KKK+K+N +QEW+ IE ++ E
Sbjct 522 QQFKSKEEYEKYIGEIEFVPKAALQYGRRMAGEF-APGGKKKRKMNIDQEWKKIEGMLKE 580
Query 78 KK 79
KK
Sbjct 581 KK 582
> xla:446708 MGC83922 protein; K13109 IK cytokine
Length=510
Score = 40.0 bits (92), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 45/76 (59%), Gaps = 3/76 (3%)
Query 7 GAPRGPTRRGPQQFASKAAYEEYMNSIELVPKAAYSYGRRIAGDINSSKNKK-KQKLNPE 65
G +GP G F ++ Y +YMN+ E +PKAA+ YG +++ + + K+ +K +
Sbjct 425 GNKKGPL--GRWDFDTQEEYSDYMNNKEALPKAAFQYGIKMSEGRKTRRFKETNEKAELD 482
Query 66 QEWRAIEKLIAEKKQL 81
++W+ I +I ++K+L
Sbjct 483 RQWKKISAIIEKRKKL 498
> hsa:3550 IK, CSA2, MGC59741, RED; IK cytokine, down-regulator
of HLA II; K13109 IK cytokine
Length=557
Score = 39.7 bits (91), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 44/76 (57%), Gaps = 3/76 (3%)
Query 7 GAPRGPTRRGPQQFASKAAYEEYMNSIELVPKAAYSYGRRIAGDINSSKNKK-KQKLNPE 65
G +GP G F ++ Y EYMN+ E +PKAA+ YG +++ + + K+ K +
Sbjct 472 GNKKGPL--GRWDFDTQEEYSEYMNNKEALPKAAFQYGIKMSEGRKTRRFKETNDKAELD 529
Query 66 QEWRAIEKLIAEKKQL 81
++W+ I +I ++K++
Sbjct 530 RQWKKISAIIEKRKKM 545
> mmu:24010 Ik, MuRED; IK cytokine; K13109 IK cytokine
Length=557
Score = 39.7 bits (91), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 44/76 (57%), Gaps = 3/76 (3%)
Query 7 GAPRGPTRRGPQQFASKAAYEEYMNSIELVPKAAYSYGRRIAGDINSSKNKK-KQKLNPE 65
G +GP G F ++ Y EYMN+ E +PKAA+ YG +++ + + K+ K +
Sbjct 472 GNKKGPL--GRWDFDTQEEYSEYMNNKEALPKAAFQYGIKMSEGRKTRRFKETNDKAELD 529
Query 66 QEWRAIEKLIAEKKQL 81
++W+ I +I ++K++
Sbjct 530 RQWKKISAIIEKRKRM 545
> dre:322241 ik, wu:fb54c03, zgc:56500; IK cytokine; K13109 IK
cytokine
Length=548
Score = 39.3 bits (90), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 22/76 (28%), Positives = 45/76 (59%), Gaps = 3/76 (3%)
Query 7 GAPRGPTRRGPQQFASKAAYEEYMNSIELVPKAAYSYGRRIAGDINSSKNKK-KQKLNPE 65
G +GP G F ++ Y +YMN+ E +PKAA+ YG +++ + + K+ +K +
Sbjct 463 GNKKGPL--GRWDFDTQEEYSDYMNNKEALPKAAFQYGIKMSEGRKTRRFKETNEKAELD 520
Query 66 QEWRAIEKLIAEKKQL 81
++W+ I +I ++K++
Sbjct 521 RQWKKISAIIEKRKKM 536
> ath:AT2G26460 RED family protein; K13109 IK cytokine
Length=585
Score = 29.6 bits (65), Expect = 2.4, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 37/76 (48%), Gaps = 6/76 (7%)
Query 6 GGAPRGPTRRGPQQFASKAAYEEYMNSIELVPKAAYSYGRRIAGDINSSKN--KKKQKLN 63
GG +G R F ++ +E+Y E +PKAA+ +G ++ + K + QKLN
Sbjct 490 GGKAKGGLHRW--DFETEEEWEKYNEQKEAMPKAAFQFGVKMQDGRKTRKQNRDRDQKLN 547
Query 64 PEQEWRAIEKLIAEKK 79
E I K++ KK
Sbjct 548 --NELHQINKILTRKK 561
> cel:Y49F6B.4 smu-2; Suppressor of Mec and Unc defects family
member (smu-2); K13109 IK cytokine
Length=547
Score = 29.3 bits (64), Expect = 3.4, Method: Compositional matrix adjust.
Identities = 20/65 (30%), Positives = 33/65 (50%), Gaps = 6/65 (9%)
Query 19 QFASKAAYEEYMNSIELVPKAAYSYGRRIAGDINSSKNKKKQKLNP----EQEWRAIEKL 74
F ++ Y YM E +PKAAY YG + KNKK+ ++ ++E I K+
Sbjct 468 DFDTEEEYASYMEGREALPKAAYQYG--VKNGEGGRKNKKQSAVSDAKRLDRELNEINKI 525
Query 75 IAEKK 79
+ ++K
Sbjct 526 MDKRK 530
> cel:Y45G5AM.7 hypothetical protein
Length=433
Score = 28.9 bits (63), Expect = 4.3, Method: Compositional matrix adjust.
Identities = 14/40 (35%), Positives = 23/40 (57%), Gaps = 0/40 (0%)
Query 28 EYMNSIELVPKAAYSYGRRIAGDINSSKNKKKQKLNPEQE 67
E IEL+ K A Y +RI DI + + + K++++ QE
Sbjct 251 ESTEKIELIQKEAEDYRKRIEEDIRNRQQRHKKEVDAHQE 290
Lambda K H
0.309 0.128 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2053886800
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40